DQBM's roundup of 2020
We are UZH's Department of Quantitative Biomedicine (DQBM)
Founded in 2019, the DQBM's mission is to foster research and education at the interface of biomedical research, bio-technology, and computational biology, to develop the foundations of next-generation precision medicine. Ultimately, our goal is to advance precision medicine for the benefit of patients.
We aim to strengthen quantitative biomedicine in Zurich
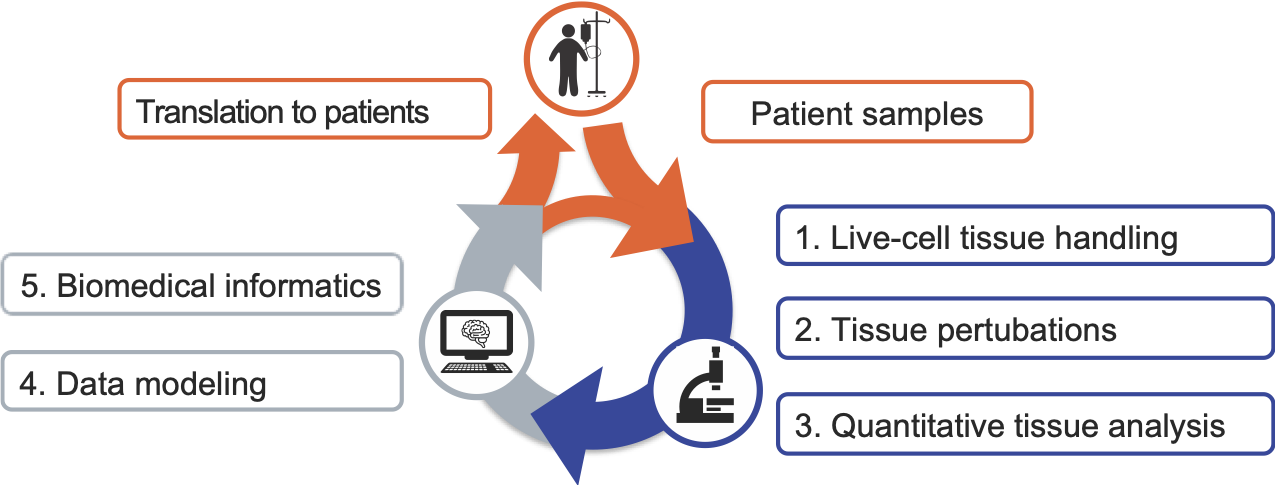
To achieve this, we take on the challenge of bridging the gap between basic research, data science and translational / clinical research groups. Specifically, we aim to strengthen quantitative biomedicine in Zurich by combining basic research and the development of novel quantitative methods for the molecular analysis of patient samples, with translational research and the development of medical informatics (Figure 1).
Ambitious goals require excellent science
To fulfil our mission, we rely on excellent science done by outstanding scientists. Below, we highlight the academic achievements of DQBM's members in 2020.
Appointments
The following professorships were established at DQBM in 2020:
- Bjoern Menze was appointed Full Professor for Biomedical Image Analysis and Machine Learning, funded by the Helmut Horten Foundation, as of 01.09.2020.
- Bernd Bodenmiller was appointed Dual Professor for Quantitative Biomedicine at UZH and ETHZ, as of 01.10.2020.
Grants and Fellowships
2020 was a successful fundraising year for DQBM's scientists. Notably:
- Bernd Bodenmiller received an SNSFproject grant to study COVID-19 immunity, in collaboration with Prof. Onur Boyman.
- Bernd Bodenmiller was supported as consortium member of the collaborative research initiative SKINTEGRITY.
- Bernd Bodenmiller received project funding from the Cancer Research Center Zurich, in collaboration with Prof. Holger Moch.
- Bernd Bodenmiller received funding as consortium member of the Hochschulmedizin Zürich Flagship project ImmunoPhage.
- Michael Krauthammer is project partner on two Innosuisse grants for clinical data science projects.
- Michael Krauthammer received funding from The LOOP Zurich.
- Michael Krauthammer will contribute his AI expertise as consortium partner of the University Research Priority Program (URPP) "Human Reproduction Reloaded".
- Magdalini Polymenidou received a competitive EU Consortium Grant from the EU Joint Programme - Neurodegenerative Disease Research (JPND) as coordinator of the ImageTDP43 consortium.
- Magdalini Polymenidou received an Innosuisse grant to evaluate the efficacy of a drug candidate for ALS.
- Magdalini Polymenidou received an SNSF project grant to explore the molecular basis of disease heterogeneity in TDP-43 proteinopathies.
- Magdalini Polymenidou received an NIH grant to study the development of immunotherapy targeting pathological TDP-43 in ALS and FTLD.
DQBM's junior scientists also did a fantastic job in securing research funds:
- Nicolas Damond (Bodenmiller lab) was awarded a JDRF FY20 postdoctoral fellowship.
- Nils Eling (Bodenmiller lab) received a Marie Skłodowska-Curie Actions Postdoctoral Fellowship to analyse heterogeneity formation in breast cancer organoids.
- Nicolas Perez (Krauthammer lab) received an UZH Entrepreneur Fellowship.
- Zsolt Balázs (Krauthammer lab) received an UZH Forschungskredit for the project “Cell-free DNA as a predictive biomarker in preeclampsia”.
- Pierre De Rossi (Polymenidou lab) was awarded the Career Development Award from Synapsis Foundation, which supports advanced postdoctoral researchers at Swiss Universities and Research Institutions seeking to establish themselves as independent researchers in the field of neurodegenerative disorders.
- Vera Wiersma (Polymenidou lab) received a FEBS Long-Term Fellowship.
Awards and Prizes
The Following DQBM members were honored for their scientific excellence:
- Magdalini Polymenidou was awarded the Franco Regli Prize 2018/2019 (2nd Prize ex æquo) for her work: 'TDP-43 extracted from frontotemporal lobar degeneration subject brains displays distinct aggregate assemblies and neurotoxic effects reflecting disease progression rates' (link).
- Johanna Wagner-Albrecht (Bodenmiller lab) was awarded the Annual Award 2020 from UZH's Faculty of Science for her PhD thesis: 'Single-Cell Proteomic Characterization of the Tumor and Immune Ecosystem of Human Breast Cancer with Focus on Metastatic Potential'.
Graduations
The following bright minds completed their PhD thesis in 2020:
- Katharina Hembach (Polymenidou lab): 'Computational analyses of high-throughput sequencing data to understand RNA metabolism defects associated with ALS'.
- Sonu Sahadevan (Polymenidou lab): 'Understanding the role of FUS in synaptic RNA homeostasis and its misregulation in neurodegenerative diseases'.
Furthermore, the following MSc students graduated in 2020 with a DQBM MSc thesis:
- Michelle Daniel (Bodenmiller lab): 'Towards characterisation of β cell heterogeneity: coupling human pancreatic islet culture and imaging mass cytometry'.
- Tobias Hoch (Bodenmiller lab): 'Characterization of chemokine expression in metastatic melanoma using simultaneous multiplexed mRNA and protein imaging by Mass Cytometry'.
- Johanna Furrer (Polymenidou lab): 'Developing new cellular models to explore TDP-43-associated pathologies'.
- Laura de Vos (Polymenidou lab): 'Development of new cellular models to study TDP-43 associated proteinopathies'.
Scientific publications
Bodenmiller lab:
Cytomapper: an R/bioconductor package for visualisation of highly multiplexed imaging data.
Eling N, Damond N, Hoch T, Bodenmiller B.
Bioinformatics. 2020 Dec 26:btaa1061.
doi: 10.1093/bioinformatics/btaa1061.
A quantitative analysis of the interplay of environment, neighborhood, and cell state in 3D spheroids.
Zanotelli VR, Leutenegger M, Lun XK, Georgi F, de Souza N, Bodenmiller B.
Mol Syst Biol. 2020 Dec;16(12):e9798.
doi: 10.15252/msb.20209798.
Mechanistic Model of Signaling Dynamics Across an Epithelial Mesenchymal Transition.
Wade JD, Lun XK, Zivanovic N, Voit EO, Bodenmiller B.
Front Physiol. 2020 Nov 30;11:579117.
doi: 10.3389/fphys.2020.579117
LifeTime and improving European healthcare through cell-based interceptive medicine.
Rajewsky N, Almouzni G, Gorski SA, Aerts S, Amit I, Bertero MG, Bock C, Bredenoord AL, Cavalli G, Chiocca S, Clevers H, De Strooper B, Eggert A, Ellenberg J, Fernández XM, Figlerowicz M, Gasser SM, Hubner N, Kjems J, Knoblich JA, Krabbe G, Lichter P, Linnarsson S, Marine JC, Marioni JC, Marti-Renom MA, Netea MG, Nickel D, Nollmann M, Novak HR, Parkinson H, Piccolo S, Pinheiro I, Pombo A, Popp C, Reik W, Roman-Roman S, Rosenstiel P, Schultze JL, Stegle O, Tanay A, Testa G, Thanos D, Theis FJ, Torres-Padilla ME, Valencia A, Vallot C, van Oudenaarden A, Vidal M, Voet T; LifeTime Community Working Groups.
Nature. 2020 Nov;587(7834):377-386.
doi: 10.1038/s41586-020-2715-9.
High-Dimensional T Helper Cell Profiling Reveals a Broad Diversity of Stably Committed Effector States and Uncovers Interlineage Relationships.
Tortola L, Jacobs A, Pohlmeier L, Obermair FJ, Ampenberger F, Bodenmiller B, Kopf M.
Immunity. 2020 Sep 15;53(3):597-613.e6.
doi: 10.1016/j.immuni.2020.07.001.
Profiling Cell Signaling Networks at Single-cell Resolution.
Lun XK, Bodenmiller B.
Mol Cell Proteomics. 2020 May;19(5):744-756.
doi: 10.1074/mcp.R119.001790
Uncovering axes of variation among single-cell cancer specimens.
Chen WS, Zivanovic N, van Dijk D, Wolf G, Bodenmiller B, Krishnaswamy S.
Nat Methods. 2020 Mar;17(3):302-310.
doi: 10.1038/s41592-019-0689-z.
The single-cell pathology landscape of breast cancer.
Jackson HW, Fischer JR, Zanotelli VRT, Ali HR, Mechera R, Soysal SD, Moch H, Muenst S, Varga Z, Weber WP, Bodenmiller B.
Nature. 2020 Feb;578(7796):615-620.
doi: 10.1038/s41586-019-1876-x.
Stabilized Reconstruction of Signaling Networks from Single-Cell Cue-Response Data.
Kumar S, Lun XK, Bodenmiller B, Rodríguez Martínez M, Koeppl H.
Sci Rep. 2020 Jan 27;10(1):1233.
doi: 10.1038/s41598-019-56444-5.
Krauthammer lab:
Differential immunomodulatory Effect of PARP Inhibition in BRCA1 deficient and competent tumor cells.
Alvarado-Cruz I, Mahmoud M, Khan M, Zhao S, Oeck S, Meas R, Clairmont K, Quintana V, Zhu Y, Porciuncula A, Wyatt H, Ma S, Shyr Y, Kong Y, LoRusso PM, Laverty D, Nagel ZD, Schalper KA,Krauthammer M, Sweasy JB.
Biochem Pharmacol. 2020 Dec 4:114359.
doi: 10.1016/j.bcp.2020.114359.
The Association of MUC16 Mutation with Tumor Mutation Burden and Its Prognostic Implications in Cutaneous Melanoma.
Wang X, Yu X, Krauthammer M, Hugo W, Duan C, Kanetsky PA, Teer JK, Thompson ZJ, Kalos D, Tsai KY, Smalley KSM, Sondak VK, Chen YA, Conejo-Garcia JR.
Cancer Epidemiol Biomarkers Prev. 2020 Sep;29(9):1792-1799.
doi: 10.1158/1055-9965.EPI-20-0307.
The Gene Expression Profile of Uropathogenic Escherichia coli in Women with Uncomplicated Urinary Tract Infections Is Recapitulated in the Mouse Model.
Frick-Cheng AE, Sintsova A, Smith SN, Krauthammer M, Eaton KA, Mobley HLT.
mBio. 2020 Aug 11;11(4):e01412-20.
doi: 10.1128/mBio.01412-20.
AutoDiscern: rating the quality of online health information with hierarchical encoder attention-based neural networks.
Kinkead L, Allam A, Krauthammer M.
BMC Med Inform Decis Mak. 2020 Jun 9;20(1):104.
doi: 10.1186/s12911-020-01131-z.
Lost in Anonymization - A Data Anonymization Reference Classification Merging Legal and Technical Considerations.
Vokinger KN, Stekhoven DJ, Krauthammer M.
J Law Med Ethics. 2020 Mar;48(1):228-231.
doi: 10.1177/1073110520917025.
Kümmerli lab:
Loss of a pyoverdine secondary receptor in Pseudomonas aeruginosa results in a fitter strain suitable for population invasion.
González J, Salvador M, Özkaya, Ö, Spick M, Reid K, Costa C, Bailey MJ, Avignone Rossa C, Kümmerli R & Jiménez JI.
ISME J 2020 Dec 15.
https://doi.org/10.1038/s41396-020-00853-2
Microbial Mutualism: Will You Still Need Me, Will You Still Feed Me?
Figueiredo ART, Kümmerli R.
Curr Biol. 2020 Sep 21;30(18):R1041-R1043.
doi: 10.1016/j.cub.2020.07.002.
Antagonistic interactions subdue inter-species green-beard cooperation in bacteria.
Sathe S, Kümmerli R.
J Evol Biol. 2020 Sep;33(9):1245-1255.
doi: 10.1111/jeb.13666.
Strain Background, Species Frequency, and Environmental Conditions Are Important in Determining Pseudomonas aeruginosa and Staphylococcus aureus Population Dynamics and Species Coexistence.
Niggli S, Kümmerli R.
Appl Environ Microbiol. 2020 Sep 1;86(18):e00962-20.
doi: 10.1128/AEM.00962-20.
Combining antibiotics with antivirulence compounds can have synergistic effects and reverse selection for antibiotic resistance in Pseudomonas aeruginosa.
Rezzoagli C, Archetti M, Mignot I, Baumgartner M, Kümmerli R.
PLoS Biol. 2020 Aug 18;18(8):e3000805.
doi: 10.1371/journal.pbio.3000805.
Competition for iron drives phytopathogen control by natural rhizosphere microbiomes.
Gu S, Wei Z, Shao Z, Friman VP, Cao K, Yang T, Kramer J, Wang X, Li M, Mei X, Xu Y, Shen Q,Kümmerli R, Jousset A.
Nat Microbiol. 2020 Aug;5(8):1002-1010.
doi: 10.1038/s41564-020-0719-8.
Bacterial siderophores in community and host interactions.
Kramer J, Özkaya Ö, Kümmerli R.
Nat Rev Microbiol. 2020 Mar;18(3):152-163.
doi: 10.1038/s41579-019-0284-4.
Harnessing bacterial interactions to manage infections: a review on the opportunistic pathogen Pseudomonas aeruginosa as a case example.
Rezzoagli C, Granato ET, Kümmerli R.
J Med Microbiol. 2020 Feb;69(2):147-161.
doi: 10.1099/jmm.0.001134.
Positive linkage between bacterial social traits reveals that homogeneous rather than specialised behavioral repertoires prevail in natural Pseudomonas communities.
Kramer J, López Carrasco MÁ, Kümmerli R.
FEMS Microbiol Ecol. 2020 Jan 1;96(1):fiz185.
doi: 10.1093/femsec/fiz185.
Menze lab:
DeepVesselNet: Vessel Segmentation, Centerline Prediction, and Bifurcation Detection in 3-D Angiographic Volumes.
Tetteh G, Efremov V, Forkert ND, Schneider M, Kirschke J, Weber B, Zimmer C, Piraud M, Menze BH.
Front Neurosci. 2020 Dec 8;14:592352.
doi: 10.3389/fnins.2020.592352.
Deep learning-enabled multi-organ segmentation in whole-body mouse scans.
Schoppe O, Pan C, Coronel J, Mai H, Rong Z, Todorov MI, Müskes A, Navarro F, Li H, Ertürk A, Menze BH.
Nat Commun. 2020 Nov 6;11(1):5626.
doi: 10.1038/s41467-020-19449-7.
Silent 3D MR sequence for quantitative and multicontrast T1 and proton density imaging.
Liu X, Gómez PA, Solana AB, Wiesinger F, Menzel MI, Menze BH.
Phys Med Biol. 2020 Sep 16;65(18):185010.
doi: 10.1088/1361-6560/aba5e8.
Spatial Distribution of Focal Lesions in Whole-Body MRI and Influence of MRI Protocol on Staging in Patients with Smoldering Multiple Myeloma According to the New SLiM-CRAB-Criteria.
Wennmann M, Hielscher T, Kintzelé L, Menze BH, Langs G, Merz M, Sauer S, Kauczor HU, Schlemmer HP, Delorme S, Goldschmidt H, Weinhold N, Hillengass J, Weber MA.
Cancers 2020 Sep 7;12(9):2537.
doi: 10.3390/cancers12092537.
Polymenidou lab:
Nuclear import receptors directly bind to arginine-rich dipeptide repeat proteins and suppress their pathological interactions
Hutten S, Usluer S, Bourgeois B, Simonetti F, Odeh HM, Fare CM, Czuppa M, Hruska-Plochan M, Hofweber M, Polymenidou M, Shorter J, Edbauer D, Madi T & Dormann D.
Cell Rep. 2020 Dec 22;33(12):108538.
DOI: 10.1016/j.celrep.2020.108538.
Phase Separation and Neurodegenerative Diseases: A Disturbance in the Force.
Zbinden A, Pérez-Berlanga M, De Rossi P, Polymenidou M.
Dev Cell. 2020 Oct 12;55(1):45-68.
doi: 10.1016/j.devcel.2020.09.014.
A Gut Feeling about C9ORF72.
Polymenidou M.
Trends Immunol. 2020 Sep;41(9):755-757.
doi: 10.1016/j.it.2020.07.012.
Outlook
We are thrilled about, and grateful for, all our members' achievements, opportunities and successes in this particularly challenging year. Moreover, we are looking forward to an exciting year ahead: In Autumn 2021, the wet labs will move into the new building on Campus Irchel (UZI5) and the DQBM will organise its first joint Block Course within the biomedicine curriculum. Furthermore, we will proclaim our founding with a celebratory DQBM Founding Symposium on 29.11.2021 and, of course, we will be doing more excellent science. Please stay tuned!