DQBM's roundup of 2019
We are UZH's Department of Quantitative Biomedicine (DQBM)
DQBM was founded on 1.1.2019. Our joint mission is to foster research and education at the interface of biomedical research, biotechnology, and computational biology, to develop the foundations of next generation precision medicine. Ultimately, our goal is to advance precision medicine for the benefit of patients.
We aim to strengthen quantitative biomedicine in Zurich
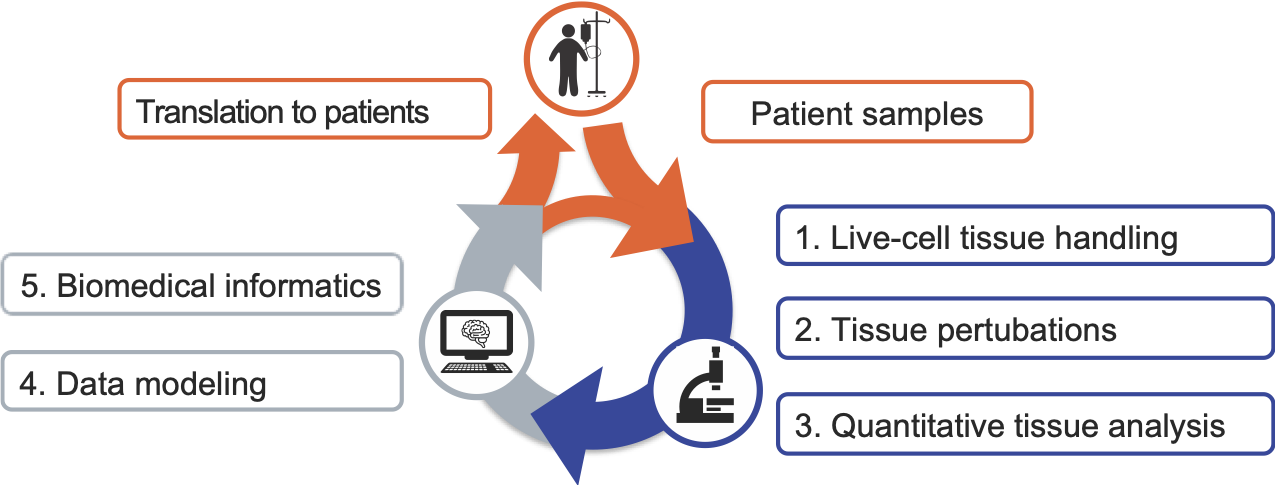
To achieve this, we take on the challenge of bridging the gap between basic research, data science and translational / clinical research groups. Specifically, we aim to strengthen quantitative biomedicine in Zurich by combining basic research and the development of novel quantitative methods for the molecular analysis of patient samples, with translational research and the development of medical informatics (Figure 1).
Ambitious goals require excellent science
To fulfil our mission, we rely on excellent science done by outstanding scientists. Below, we highlight the academic achievements of DQBM's members in 2019.
Appointments
The following professorships were established at DQBM in 2019:
- Bernd Bodenmiller was appointed Associate Professor for Quantitative Cell Biology, as of 1 March 2019.
- Magdalini Polymenidou was appointed Associate Professor for Biomedicine,
in particular Molecular Pathogenesis of Neurodegeneration, as of 1 October 2019.
Grants and Fellowships
2019 was a great fundraising year for DQBM's scientists. Notably:
- Bernd Bodenmiller received a coveted ERC Consolidator Grant to analyze functional tissue motifs for precision medicine in metastatic breast cancer.
- Bernd Bodenmiller received a IMI2 Human Tumour Microenvironment Immunoprofiling Grant.
- Rolf Kümmerli received a Novartis Foundation grant for medical-biological research.
- Michael Krauthammer became project partner on the Infectious Disease Biobank Zurich Project, funded by SNF.
- Magdalini Polymenidou received an FTD Biomarker Initiative grant from AFTD to develop biomarkers for FTD patients.
DQBM's junior scientists also did a fantastic job in securing research funds:
- Hartland Jackson (Bodenmiller lab) received a Scholarship for the Next Generation of Scientists from the Cancer Research Society.
- Merrick Strotton (Bodenmiller lab) was awarded an SNF SPARK grant to develop new multiplex imaging approaches.
- Nils Eling (Bodenmiller lab) received an EMBO Postdoctoral Fellowship to analyze heterogeneity formation in breast cancer organoids.
- Özhan Özkaya (Kümmerli lab) received an UZH Forschungskredit Postdoc grant to study the ecological and evolutionary consequences of cell death on public goods cooperation in bacteria using single-cell tracking.
- Xiaokang Lun (Bodenmiller lab) received an SNF Early Postdoc.Mobility Fellowship.
Awards and Prizes
A number of DQBM members were honored for their scientific excellence:
- Bernd Bodenmiller received the Friedrich-Miescher-Award for outstanding research in the field of biochemistry.
- Bernd Bodenmiller was selected for the Analytical Scientist’s 2019 Power List Top 100.
- Julien Weber (Polymenidou lab) was voted Best Lab Manager of 2019 by Proteintech.
- Selina Niggli (Kümmerli lab) won the best oral presentation award on the topic Environmental Microbiology at the Annual Swiss Society for Microbiology Meeting 2019.
Graduations
The following bright minds completed their PhD thesis in 2019:
- Chiara Rezzoagli (Kümmerli lab): 'Bacterial cooperation and the design of evolutionary robust antimicrobials'.
- Johanna Wagner *with distinction* (Bodenmiller lab): 'Single-Cell Proteomic Characterization of the Tumor and Immune Ecosystem of Human Breast Cancer with Focus on Metastatic Potential'.
- Marco Tognetti (Bodenmiller lab): 'Deciphering the signaling network landscape of breast cancer supports personalized medicine'.
- Mélanie Jambeau (Polymenidou lab): 'Understanding and Targeting Dipeptide Repeat Protein Toxicity by Immunotherapy in ALS and FTD'.
- Vito Zanotelli (Bodenmiller lab): 'Investigation of Intra- and Intercellular Signaling through Mass Cytometry Based Single Cell Methods'.
- Xiaokang Lun *with distinction* (Bodenmiller lab): 'Single-cell analysis reveals protein abundance-dependent signaling network modulations'.
Furthermore, the following MSc students graduated in 2019 with a DQBM MSc thesis:
- Martina Archetti (Kümmerli lab): 'Combining antibiotics and antivirulence compounds against Pseudomonas aeruginosa PAO1'.
- Nadine Koch (Kümmerli lab): 'Polymicrobial interactions in human serum and in the insect host Galleria mellonella'.
Scientific publications
Bodenmiller lab:
Hair eruption initiates and commensal skin microbiota aggravate adverse events of anti-EGFR therapy.
Klufa J, Bauer T, Hanson B, Herbold C, Starkl P, Lichtenberger B, Srutkova D, Schulz D, Vujic I, Mohr T, Rappersberger K, Bodenmiller B, Kozakova H, Knapp S, Loy A, Sibilia M. Sci Transl Med. 2019 Dec 11;11(522). pii: eaax2693.
doi: 10.1126/scitranslmed.aax2693.
Modeling Cell-Cell Interactions from Spatial Molecular Data with Spatial Variance Component Analysis.
Arnol D, Schapiro D, Bodenmiller B, Saez-Rodriguez J, Stegle O.
Cell Rep. 2019 Oct 1;29(1):202-211.e6.
doi: 10.1016/j.celrep.2019.08.077.
Analysis of the Human Kinome and Phosphatome by Mass Cytometry Reveals Overexpression-Induced Effects on Cancer-Related Signaling.
Lun XK, Szklarczyk D, Gábor A, Dobberstein N, Zanotelli VRT, Saez-Rodriguez J, von Mering C, Bodenmiller B.
Mol Cell. 2019 Jun 6;74(5):1086-1102.e5.
doi: 10.1016/j.molcel.2019.04.021.
A Single-Cell Atlas of the Tumor and Immune Ecosystem of Human Breast Cancer.
Wagner J, Rapsomaniki MA, Chevrier S, Anzeneder T, Langwieder C, Dykgers A, Rees M, Ramaswamy A, Muenst S, Soysal SD, Jacobs A, Windhager J, Silina K, van den Broek M, Dedes KJ, Rodríguez Martínez M, Weber WP, Bodenmiller B.
Cell. 2019 May 16;177(5):1330-1345.e18.
doi: 10.1016/j.cell.2019.03.005.
IL-8 and CXCR1 expression is associated with cancer stem cell-like properties of clear cell renal cancer.
Corrò C, Healy ME, Engler S, Bodenmiller B, Li Z, Schraml P, Weber A, Frew IJ, Rechsteiner M, Moch H. J Pathol. 2019 Jul;248(3):377-389.
doi: 10.1002/path.5267.
In-Depth Characterization of Monocyte-Derived Macrophages using a Mass Cytometry-Based Phagocytosis Assay.
Schulz D, Severin Y, Zanotelli VRT, Bodenmiller B.
Sci Rep. 2019 Feb 13;9(1):1925.
doi: 10.1038/s41598-018-38127-9.
A Map of Human Type 1 Diabetes Progression by Imaging Mass Cytometry.
Damond N, Engler S, Zanotelli VRT, Schapiro D, Wasserfall CH, Kusmartseva I, Nick HS, Thorel F, Herrera PL, Atkinson MA, Bodenmiller B.
Cell Metab. 2019 Mar 5;29(3):755-768.e5.
doi: 10.1016/j.cmet.2018.11.014.
Krauthammer lab:
Neural networks versus Logistic regression for 30 days all-cause readmission prediction.
Allam A, Nagy M, Thoma G, Krauthammer M.
Sci Rep. 2019 Jun 26;9(1):9277.
doi: 10.1038/s41598-019-45685-z.
A novel anti-melanoma SRC-family kinase inhibitor.
Halaban R, Bacchiocchi A, Straub R, Cao J, Sznol M, Narayan D, Allam A, Krauthammer M, Mansour TS.
Oncotarget. 2019 Mar 19;10(23):2237-2251.
doi: 10.18632/oncotarget.26787.
Kümmerli lab:
Positive linkage between bacterial social traits reveals that homogeneous rather than specialized behavioral repertoires prevail in natural Pseudomonas communities.
Kramer J, López Carrasco MÁ, Kümmerli R.
FEMS Microbiol Ecol. 2019 Nov 26. pii: fiz185.
doi: 10.1093/femsec/fiz185.
Bacterial siderophores in community and host interactions.
Kramer J, Özkaya Ö, Kümmerli R.
Nat Rev Microbiol. 2019 Nov 20.
doi: 10.1038/s41579-019-0284-4.
Transposable temperate phages promote the evolution of divergent social strategies in Pseudomonas aeruginosa populations.
O'Brien S, Kümmerli R, Paterson S, Winstanley C, Brockhurst MA.
Proc Biol Sci. 2019 Oct 9;286(1912):20191794.
doi: 10.1098/rspb.2019.1794.
Genetic architecture constrains exploitation of siderophore cooperation in the bacterium Burkholderia cenocepacia.
Sathe S, Mathew A, Agnoli K, Eberl L, Kümmerli R. Evolution Letters.
doi:10.1002/evl3.144
In-vivo microscopy reveals the impact of Pseudomonas aeruginosa social interactions on host colonization.
Rezzoagli C, Granato ET, Kümmerli R.
ISME J. 2019 Oct;13(10):2403-2414.
doi: 10.1038/s41396-019-0442-8.
Individual- versus group-optimality in the production of secreted bacterial compounds.
Schiessl KT, Ross-Gillespie A, Cornforth DM, Weigert M, Bigosch C, Brown SP, Ackermann M, Kümmerli R.
Evolution. 2019 Apr;73(4):675-688.
doi: 10.1111/evo.13701.
Understanding policing as a mechanism of cheater control in cooperating bacteria.
Wechsler T, Kümmerli R, Dobay A.
J Evol Biol. 2019 May;32(5):412-424.
doi: 10.1111/jeb.13423.
Polymenidou lab:
SarkoSpin: A Technique for Biochemical Isolation and Characterization of Pathological TDP-43 Aggregates.
Pérez-Berlanga M, Laferrière F, Polymenidou M. Bio-protocol.
DOI:10.21769/BioProtoc.3424.
Structural Transition, Function and Dysfunction of TDP-43 in Neurodegenerative Diseases.
Afroz T, Pérez-Berlanga M, Polymenidou M. Chimia (Aarau). 2019 May 29;73(6):380-390.
doi: 10.2533/chimia.2019.380.
The Solution Structure of FUS Bound to RNA Reveals a Bipartite Mode of RNA Recognition with Both Sequence and Shape Specificity.
Loughlin FE, Lukavsky PJ, Kazeeva T, Reber S, Hock EM, Colombo M, Von Schroetter C, Pauli P, Cléry A, Mühlemann O, Polymenidou M, Ruepp MD, Allain FH. Mol Cell. 2019 Feb 7;73(3):490-504.e6.
doi: 10.1016/j.molcel.2018.11.012.
TDP-43 extracted from frontotemporal lobar degeneration subject brains displays distinct aggregate assemblies and neurotoxic effects reflecting disease progression rates.
Laferrière F, Maniecka Z, Pérez-Berlanga M, Hruska-Plochan M, Gilhespy L, Hock EM, Wagner U, Afroz T, Boersema PJ, Barmettler G, Foti SC, Asi YT, Isaacs AM, Al-Amoudi A, Lewis A, Stahlberg H, Ravits J, De Giorgi F, Ichas F, Bezard E, Picotti P, Lashley T, Polymenidou M. Nat Neurosci. 2019 Jan;22(1):65-77.
doi: 10.1038/s41593-018-0294-y.
Memory Decline and Its Reversal in Aging and Neurodegeneration Involve miR-183/96/182 Biogenesis.
Jawaid A, Woldemichael BT, Kremer EA, Laferriere F, Gaur N, Afroz T, Polymenidou M, Mansuy IM. Mol Neurobiol. 2019 May;56(5):3451-3462.
doi: 10.1007/s12035-018-1314-3.
Outlook
It is safe to say that DQBM is off to a great start. Our prognosis is that 2020 will be even more exciting, as i) we aim to expand our Faculty with a Professorship for Machine Learning in Precision Medicine, ii) we will proclaim our founding with a celebratory DQBM Founding Symposium on 19 October 2020, and iii) we will be doing more excellent science. Please stay tuned!